Collaboration is key: new cross-discipline approaches to glioma – interview with Christian Naus
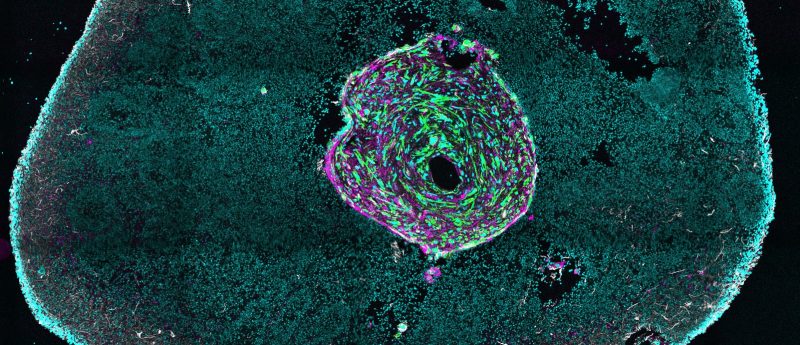
Glioma has one of the poorest cancer prognoses – a statistic that several research groups across the globe are working to change. One key approach is the development of suitable models for assessing the efficacy of novel therapeutics; something that Christian Naus, Professor at The University of British Columbia (BC, Canada) and Canada Research Chair in Gap Junctions in Neurological Disorders, is currently working on. But with a challenge on this scale, collaboration is key: Christian is working with colleagues across disciplines and drawing on additive manufacturing expertise to develop 3D glioma models with the hope that these 3D organoids...